LDDMM: how to estimate a deterministic atlas?#
This is analog to a registration problem for multiple meshes.
For registration between two meshes check LDDMM: how to register a mesh to a template?.
[1]:
import shutil
import string
import numpy as np
import pyvista as pv
import polpo.utils as putils
from polpo.mesh.deformetrica import FrechetMean, LddmmMetric, Point
from polpo.mesh.generation.blob import create_blob
from polpo.preprocessing.mesh.registration import RigidAlignment
[KeOps] Warning : CUDA was detected, but driver API could not be initialized. Switching to CPU only.
[2]:
RECOMPUTE = False
# NB: fix seed before setting it to False
np.random.seed(42)
STATIC_VIZ = True
if STATIC_VIZ:
pv.set_jupyter_backend("static")
[3]:
OUTPUTS_DIR = putils.get_results_path() / "deterministic_atlas_blob_example"
if OUTPUTS_DIR.exists() and RECOMPUTE:
shutil.rmtree(OUTPUTS_DIR)
OUTPUTS_DIR.mkdir(exist_ok=True, parents=True)
Generate meshes#
[4]:
n_meshes = 3
bump_amp = 0.2
raw_meshes = [
create_blob(resolution=10, bump_amp=bump_amp, n_bumps=5, smoothing_iter=10)
for _ in range(n_meshes)
]
raw_meshes[0].points.shape
[4]:
(82, 3)
[5]:
pl = pv.Plotter(border=False)
for mesh in raw_meshes:
pl.add_mesh(mesh, show_edges=True, opacity=0.5)
pl.show()
[6]:
prep_pipe = RigidAlignment(known_correspondences=True)
meshes = prep_pipe(raw_meshes)
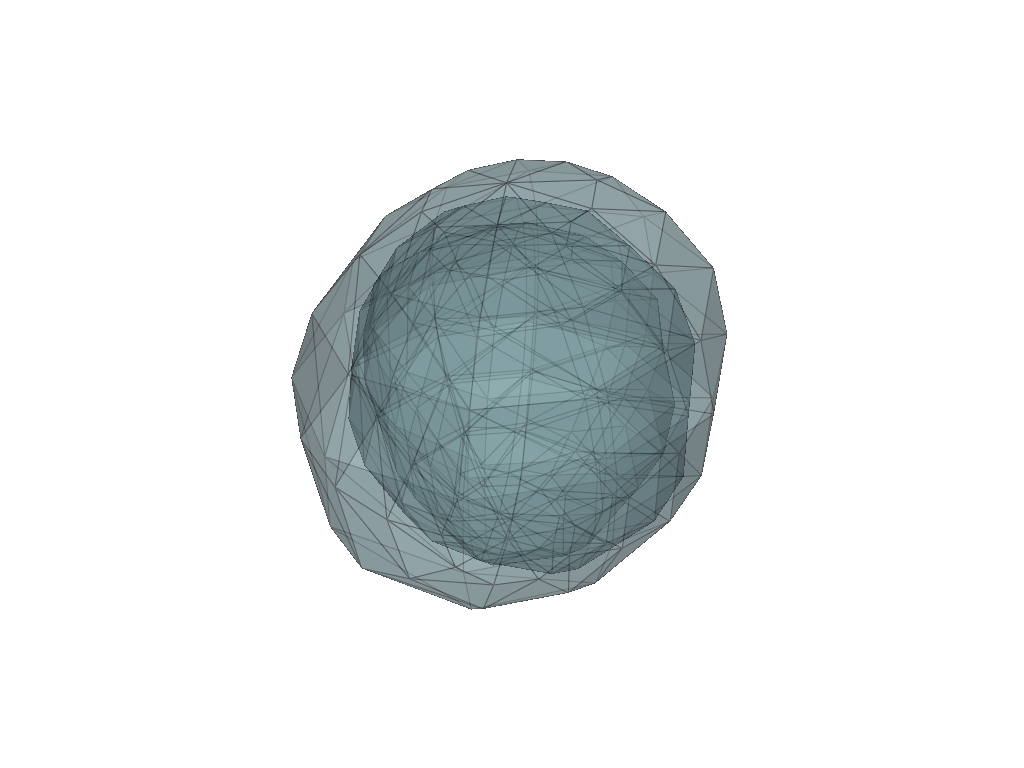
[7]:
pl = pv.Plotter(border=False)
for mesh in meshes:
pl.add_mesh(mesh, show_edges=True, opacity=0.5)
pl.show()

Deterministic atlas#
[8]:
kernel_width = 2 * bump_amp
registration_kwargs = dict(
kernel_width=kernel_width,
regularisation=1.0,
max_iter=2000,
freeze_control_points=False,
metric="varifold",
tol=1e-16,
attachment_kernel_width=bump_amp,
)
metric = LddmmMetric(OUTPUTS_DIR, **registration_kwargs)
[9]:
dataset = [
Point(id_=string.ascii_uppercase[index], pv_surface=mesh, dirname=metric.meshes_dir)
for index, mesh in enumerate(meshes)
]
Use LDDMM to estimate the atlas.
[10]:
estimator = FrechetMean(
metric,
initial_step_size=1e-1,
)
estimator.fit(dataset, atlas_id="atlas")
[10]:
<polpo.mesh.deformetrica.FrechetMean at 0x768e092eb890>
[11]:
atlas = estimator.estimate_
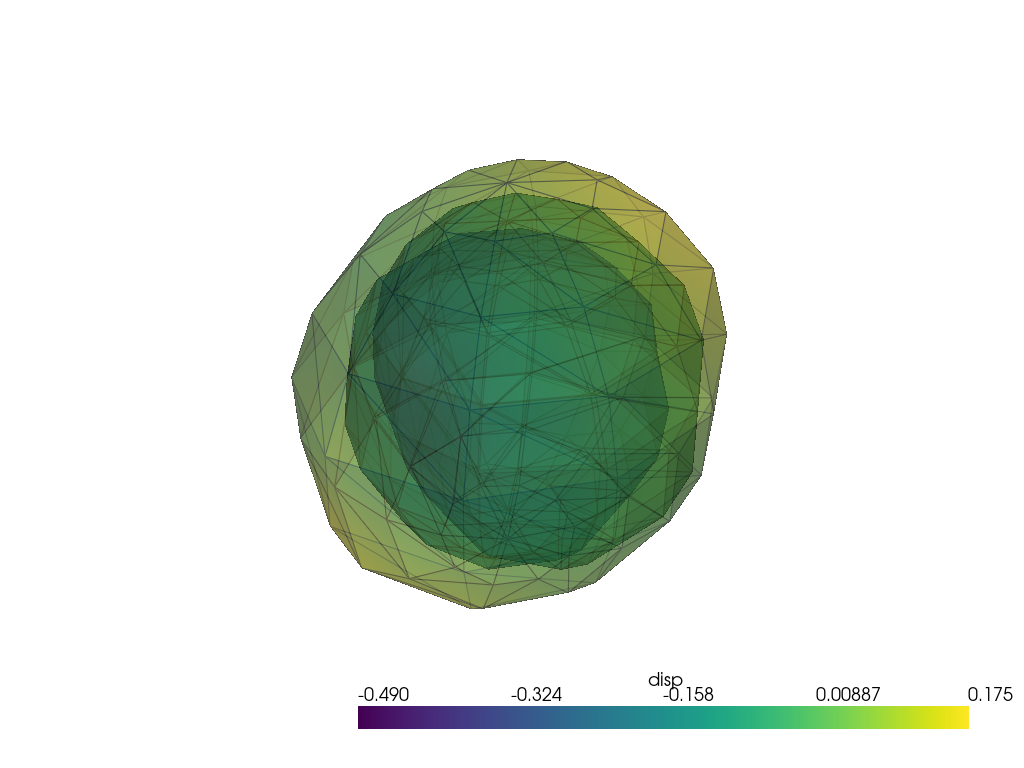
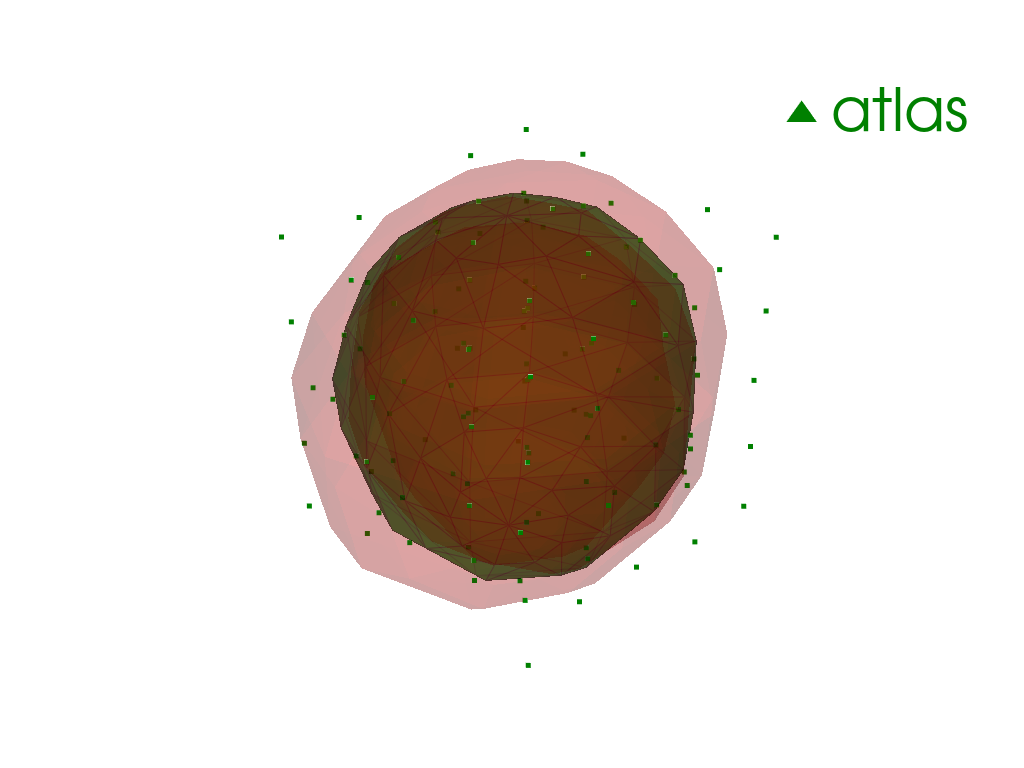
[12]:
pl = pv.Plotter(border=False)
for point in atlas.points:
pl.add_mesh(point.as_pv(), opacity=0.2, color="red")
pl.add_mesh(
atlas.as_pv(),
show_edges=True,
opacity=0.5,
color="green",
label="atlas",
)
pl.add_points(pv.PolyData(atlas.control_points(as_path=False)), color="green")
pl.add_legend()
pl.show()

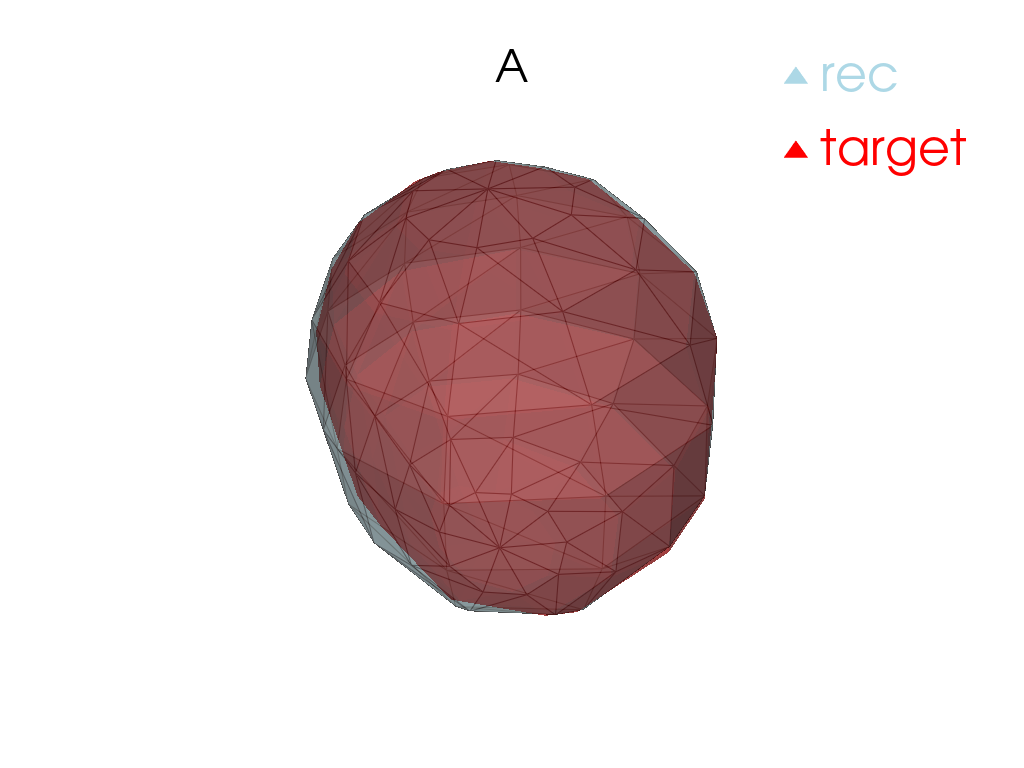
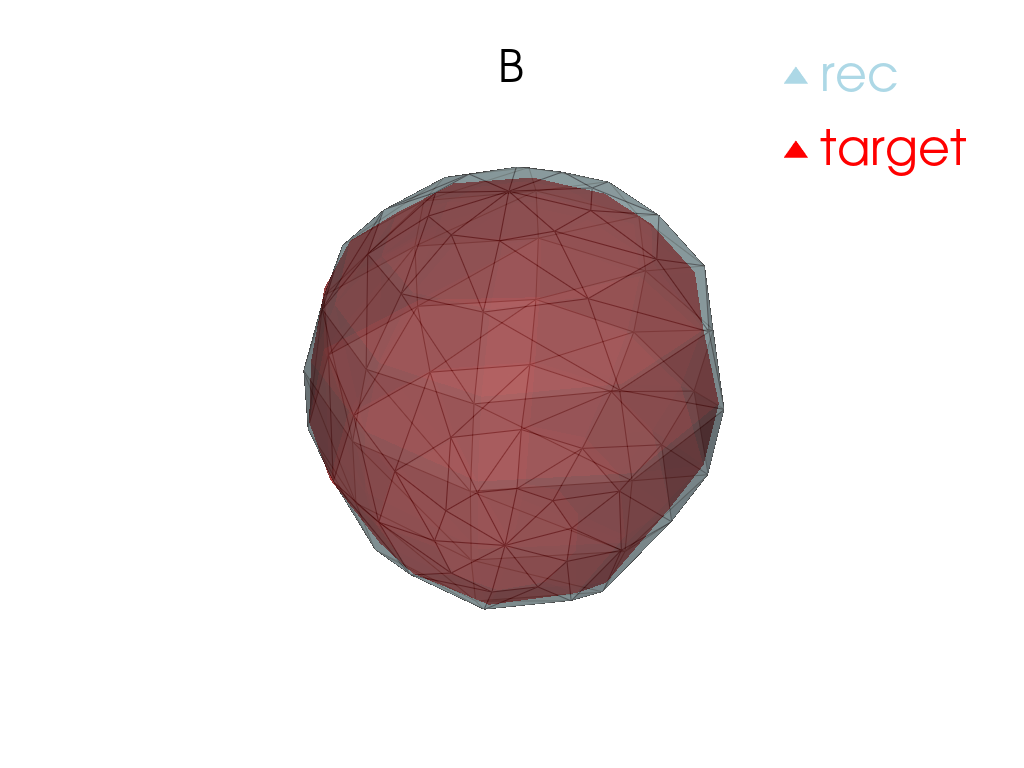
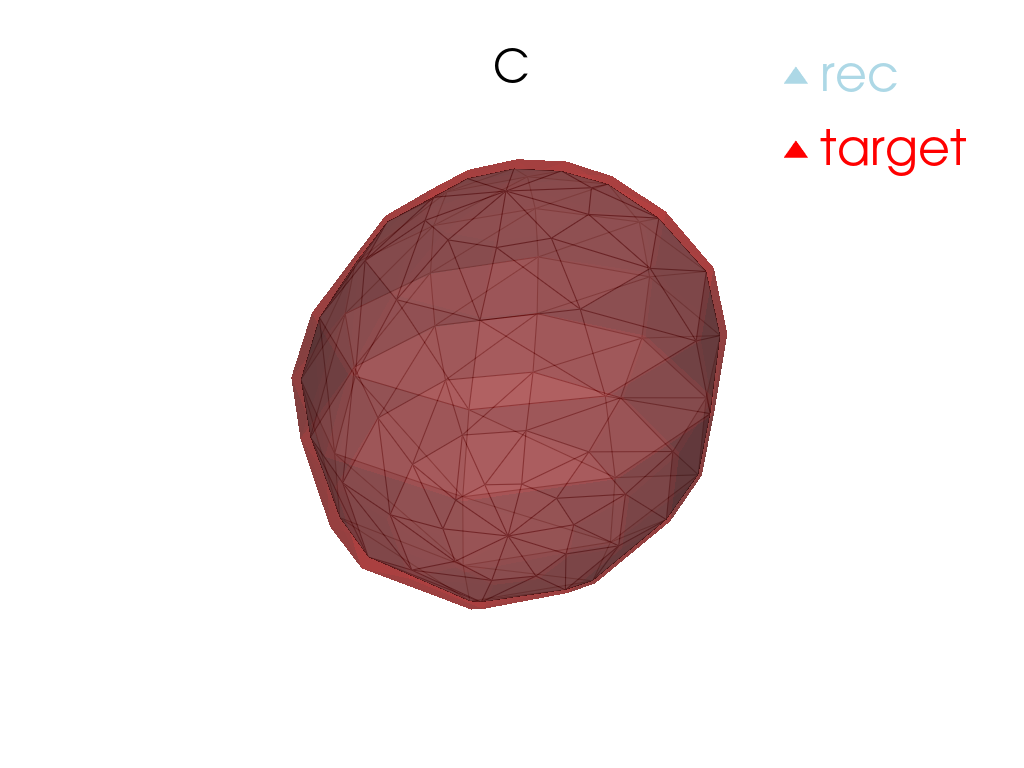
Reconstruction
[13]:
for tangent_vec in atlas.tangent_vecs():
pl = pv.Plotter()
pl.add_mesh(
tangent_vec.reconstructed().as_pv(),
show_edges=True,
opacity=0.5,
label="rec",
)
pl.add_mesh(
tangent_vec.point.as_pv(),
opacity=0.5,
color="red",
label="target",
)
pl.add_title(tangent_vec.point.id)
pl.add_legend()
pl.show()